Probe development and labeling
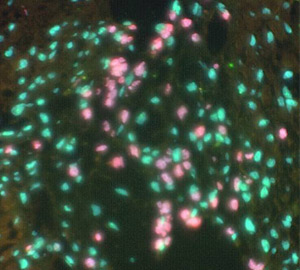
Custom pig whole-genomic paint (green) and baboon whole-genomic paint (pink)
As researchers attempt to identify specific gene loci and determine their involvement in the progression of disease, the Cytogenetics Core can develop and produce DNA, RNA and cDNA fluorescence in situ hybridization probes from various species.
Probe size can be a limiting factor in detecting genetic abnormalities. As a result, a variety of fluorophores, labeling methods and amplification techniques are available.
Details
Probe development
The core will perform internet and literature searches investigating bacterial artificial chromosome and P1-derived artificial chromosome clones for specific gene loci or sequences in which the fluorescence in situ hybridization probe will span. If the:
- Bacterial artificial chromosome or P1-derived artificial chromosome clone is available from a commercial source, the Cytogenetics Core will obtain it.
- Clone is available from a private source, the investigator is asked to obtain and ship it to the core.
Alternatively, the core will design primers for a specific DNA sequence, with primers ordered from a commercial source. In addition to a specific gene region, bacterial artificial chromosome and P1-derived artificial chromosome clones contain a significant amount of excess sequence.
Whole-genomic DNA from any species can be isolated and labeled. This type of probe is often used to determine integration in xenograft specimens.
Isolation techniques
Isolation is done from bacterial artificial chromosomes, P1-derived artificial chromosomes and whole-genomic DNA.
DNA labeling
Available are direct (nick translation and PCR) and indirect (random prime labeling and PCR) methods with a fluorescent dye for fluorescence in situ hybridization analysis.
Detection and amplification
Probes labeled in biotin or digoxigenin are detected with a secondary antibody conjugated with a red or green fluor. Alternatively, biotin-labeled probes can be detected using tyramide signal amplification conjugated to a red or green fluor to amplify the signal.
Quality control
Procedures include pre-analytical processing and confirmation by fluorescence in situ hybridization chromosome analysis. Importantly, this service will be expanded and modified to best meet the needs of the investigator.