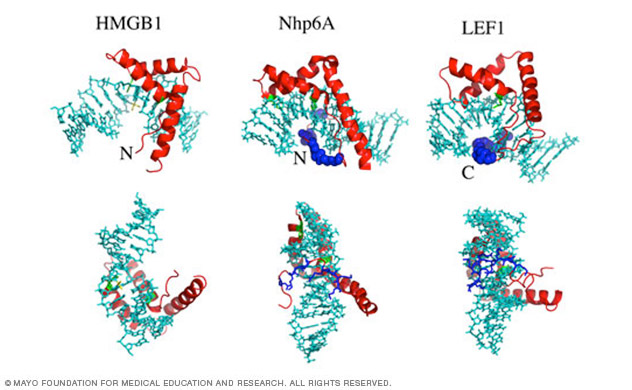
DNA bending by architectural proteins
DNA flexibility in cells
The Nucleic Acid Structure and Recognition Lab is studying the concept that the exceptional stiffness of double-stranded DNA is related to DNA charge. This concept remains controversial even 60 years after the elucidation of DNA structure. The Maher lab has shown that asymmetric charge neutralization along the DNA backbone induces DNA bending toward the neutralized face. Similar effects are seen for tethered ions and bound proteins with modified charge. Ligase-catalyzed cyclization kinetics studies have explored the flexibility of DNAs with non-natural charge densities. Results of those studies support and extend the Manning hypothesis that DNA charge density influences flexibility.
The Maher laboratory also studies the properties and mechanisms of proteins that enhance the apparent flexibility of DNA by sequence-nonspecific DNA binding and kinking. Eukaryotic high mobility group box (HMGB) proteins and bacterial Escherichia coli HU protein serve as models. Collaborative studies using single-molecule approaches and theory are revealing the DNA binding and bending properties of these proteins. The laboratory has extended collaborative single-molecule studies to understand chromatin unwinding under force, in the presence of histone chaperones, and when nucleosomes are modified by acylation.
The Maher laboratory reestablished an experimentally tractable implementation of the classic Lac repressor-mediated DNA-looping model in living bacterial cells. This work has allowed an understanding of why the "softness" of DNA appears greater in vivo. Studies are probing the importance of bacterial nucleoid proteins and supercoiling in modifying apparent DNA flexibility. These experiments help us understand how cells use proteins to assist in managing the local rigidity of DNA. This work inspired collaborative experiments to engineer new DNA-looping proteins based on designed sequence-specific transcription activator-like effector proteins.
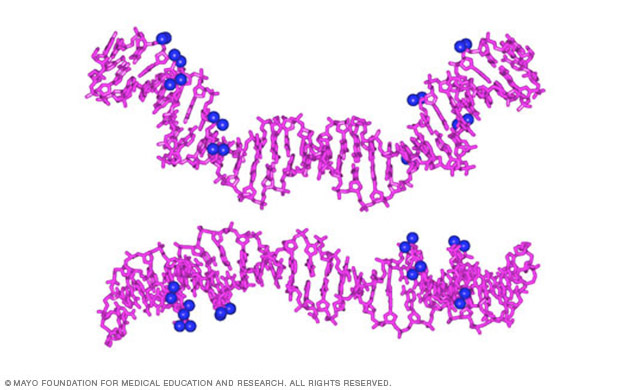
Studies of phosphate neutralization on DNA shape