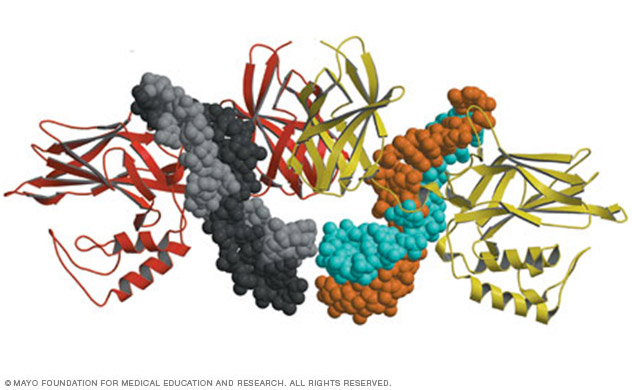
X-ray crystal structure of a novel complex between NF-kappaB protein (ribbons) and two folded RNA molecules (spheres).
Artificial gene regulation by RNA molecules
It is increasingly clear that small RNA molecules play roles in natural gene regulation. Can new examples be engineered? The Nucleic Acid Structure and Recognition Laboratory seeks to use RNA as the raw material for new kinds of gene regulators. These act by inhibiting or redirecting transcription factors.
Taking a cue from certain natural RNA decoys for DNA-binding proteins, the Maher laboratory is using in vitro selection to find and characterize RNA inhibitors. The inhibitors being implemented are from the nuclear factor kappa-light-chain-enhancer of activated B cells (NF-kappaB) family of DNA-binding transcription factors. NF-kappaB proteins are key regulators of pathological inflammation, HIV-1 viral activation, and tumor cell resistance to apoptosis after chemotherapy and radiation.
In vitro, yeast genetic, structural biology and model organism approaches are being used to test the potential of such decoy RNA molecules. Strategies for therapeutic viral expression of such RNAs in target cells are being explored. Yeast genetic selections also are being applied to select RNAs from random sequence libraries that can redirect transcription activators and repressors to control genes of interest in disease therapy and genetic engineering.