-

Discovering new therapeutic treatments for inflammatory bowel disease
Dr. Faubion's lab seeks to discover, identify and develop individualized treatments to improve treatment options for people with inflammatory bowel disease.
-
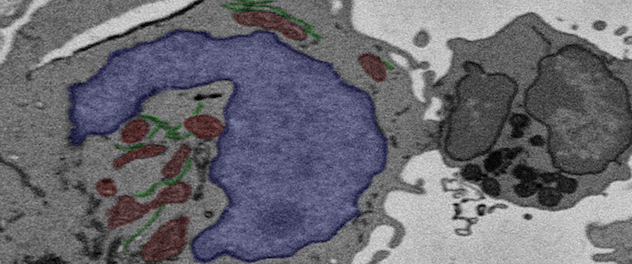
Characterizing the epigenetics of T-cell fate decisions
Understanding the regulatory mechanisms necessary for T-regulatory (Treg) gene expression will help improve Treg suppression in the setting of intestinal inflammation.
-
Understanding inflammatory and epigenetic mechanisms driving inflammatory bowel disease pathophysiology
Dr. Faubion's lab uses colitis mouse models and humanized mouse models to understand the loss of immunosuppressive function in T-regulatory cells.
-
Building an inflammatory bowel disease biobank to discover molecular signatures
Clinical samples are a critical component in the lab's research to discover new therapeutic targets by examining the molecular signatures responsible for clinical phenotypes. Dr. Faubion's team has an active biobank of inflammatory bowel disease samples to analyze using discovery platforms.
-
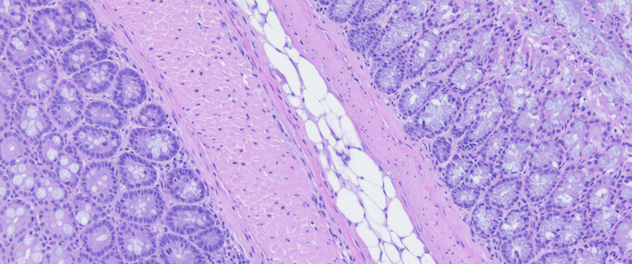
Using a systems biology approach to study inflammatory bowel disease pathophysiology
By using stool, biopsies, serum and immune cells, the lab takes a systems biology approach toward understanding how molecular networks influence chronic intestinal inflammation.
Overview
The Immuno-Epigenetics Laboratory led by William A. Faubion Jr., M.D., focuses on defining how immune cell identity is established, maintained and dysregulated in inflammatory bowel disease (IBD). The lab's work centers on the premise that immune function is not fixed. Rather, it's encoded through epigenetic programs that integrate environmental, metabolic and inflammatory signals to determine cell state.
The lab studies the epigenetic mechanisms that govern T-cell fate, particularly T-regulatory cells (Tregs) and effector populations, and how chronic intestinal inflammation alters these programs. The long-term objective is not simply to describe these states but to make them actionable. The lab seeks to identify mechanisms that can be manipulated to restore immune tolerance.
The lab's approach integrates human tissue, high-dimensional molecular profiling and functional modeling systems to define disease-relevant immune states and the regulatory circuitry that sustains them. These efforts are directly linked to therapeutic development, including engineered cell therapies and epigenetic-targeted interventions.
The lab's approach
The Immuno-Epigenetics Lab takes a mechanistic and translational approach to immunology. The lab:
- Defines pathogenic and regulatory immune cell states in human IBD.
- Identifies epigenetic regulators that control T-cell differentiation, stability and function.
- Resolves how metabolic and environmental signals are encoded into chromatin state.
- Engineers regulatory T cells ex vivo to enhance suppressive function.
- Translates these findings into early-phase clinical trials.
Translational focus
A central priority of the lab is to move beyond association toward intervention. The lab has translated multiple discoveries into clinical applications, including completing early-phase trials and ongoing phase 2 studies. These efforts focus on modulating immune cell behavior through epigenetic reprogramming or cellular engineering to restore durable immune regulation in patients with IBD.
This bidirectional framework — mechanism informing therapy and clinical observation refining mechanism — drives the lab's overall structure.
Research strategy
The lab is organized around a set of linked mechanistic and translational objectives:
- Define the epigenetic circuits governing Treg and TH17 cell fate and plasticity.
- Identify metabolically coupled pathways that reinforce pathogenic immune states.
- Dissect chromatin-based mechanisms, such as polycomb complexes, G9a or BMI1, that stabilize or destabilize immune identity.
- Develop ex vivo strategies to optimize Treg function for adoptive cell therapy.
- Target epigenetic regulators with small molecules to reprogram immune responses.
- Integrate multi-omic datasets to identify tractable therapeutic nodes.
Where the field is going
The field of molecular immunology is shifting from static classification of immune cells toward a model of dynamic cell state. In IBD, this has several implications:
- Pathogenicity reflects unstable or misprogrammed immune states rather than fixed lineages.
- Epigenetic regulators function as control points that can be therapeutically targeted.
- Single-cell and spatial technologies allow direct resolution of these states in human tissue.
- Cell-based therapies, particularly engineered Tregs, represent a viable strategy for restoring immune tolerance.
The lab's work is positioned at this interface: defining these states mechanistically and developing strategies to manipulate them in a clinically meaningful way.