Research projects
Our laboratory investigates the biological and metabolic mechanisms that enable brain tumors, particularly glioblastoma, to resist therapy and evade immune control.
By integrating tumor biology, immunology and metabolic engineering, our research projects aim to translate fundamental discoveries into next-generation immunotherapeutic strategies for adults and children with brain tumors.
Metabolic interactions in the glioblastoma tumor microenvironment
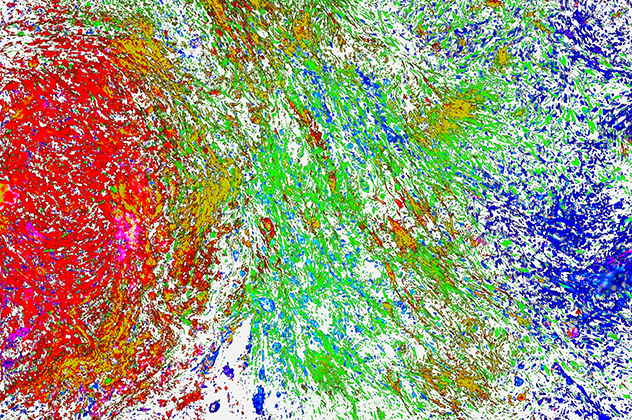 Spatial complexity of the glioblastoma tumor microenvironment
Spatial complexity of the glioblastoma tumor microenvironment
High-resolution fluorescent imaging of the glioblastoma tumor microenvironment illustrates its profound cellular heterogeneity. Distinct colors represent different cell populations, revealing nonrandom spatial organization, variable cell density, and coordinated patterns of distribution and migration that reflect structured interactions within the tumor ecosystem. These influence tumor progression, immune suppression and therapeutic response.
Glioblastoma develops within a highly heterogeneous and immunosuppressive tumor microenvironment that actively promotes therapy resistance.
A central focus of our lab is to dissect how tumor cells communicate with immune populations, particularly tumor-associated macrophages and other immune-suppressive cells, through metabolic signaling and resource exchange. Our work examines how these interactions support the survival of treatment-resistant tumor cell states and impair effective antitumor immunity.
By identifying the metabolic dependencies and signaling pathways that sustain this pathological crosstalk, we aim to uncover actionable vulnerabilities that can be therapeutically exploited.
Our long-term objective is to disrupt tumor-supportive immune niches and translate these insights into clinically relevant strategies that improve outcomes for people with glioblastoma.
Immunotherapeutic targeting of slow-cycling cancer stem cells in glioblastoma
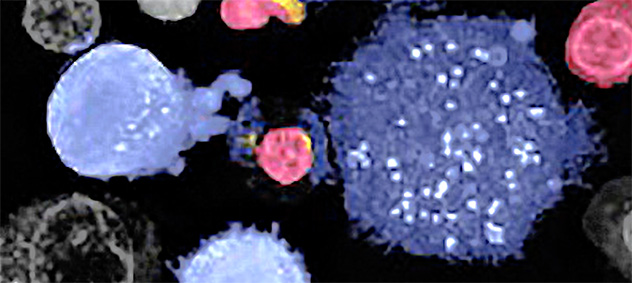 Engineered T-cell engagement of cancer stem cells
Engineered T-cell engagement of cancer stem cells
Holotomography-based time-lapse imaging shows engineered T cells (smaller cells) cocultured with brain cancer stem cells. The image captures a primed T cell engaging and attacking two cancer stem cells, which begin to exhibit morphological changes consistent with cell death, illustrating the dynamic and specific nature of immune-mediated tumor targeting.
Glioblastoma recurrence is driven in part by rare tumor cell populations that tolerate conventional therapies and reinitiate tumor growth.
Our laboratory identified a distinct pool of slow-cycling, highly tumorigenic cancer stemlike cells that exhibit pronounced resistance to radiation and chemotherapy.
This research project leverages the ability to isolate and molecularly characterize these clinically relevant cell populations to develop a novel immunotherapy strategy.
By extracting immunogenic RNA signatures from slow-cycling tumor cells, we aim to educate and activate immune effector cells, particularly T cells, to recognize and eliminate the cellular drivers of recurrence.
This approach seeks to move beyond bulk tumor targeting and instead focus immune pressure on the most dangerous tumor cell states, with the ultimate goal of preventing disease relapse.
Metabolic engineering of T cells for enhanced antitumor activity
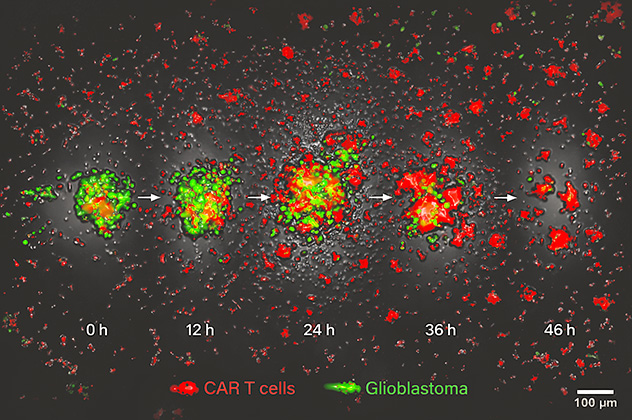 Temporal dynamics of CAR-T cell-mediated glioblastoma elimination
Temporal dynamics of CAR-T cell-mediated glioblastoma elimination
Time-lapse imaging shows metabolically engineered CAR-T cells (red) targeting glioblastoma cells (green). The series demonstrates how metabolic optimization enables T-cell engagement, effective tumor clearance and eventual immune dominance within the culture, modeling strategies to overcome tumor-driven metabolic suppression.
Adoptive T-cell therapies represent a promising avenue for treating brain tumors. However, their efficacy is frequently limited by the extreme metabolic and immunosuppressive constraints of the tumor microenvironment.
Tumor cells aggressively compete for essential nutrients, such as glucose, which impairs T-cell activation, persistence and effector function.
In this research project, we're investigating strategies to metabolically reprogram antitumor T cells to enhance their fitness and functionality in nutrient-restricted environments.
Our approaches include genetic modification of metabolic pathways, in vitro metabolic conditioning, and validation in patient-derived and in vivo brain tumor models.
By aligning immune cell metabolism with the demands of the tumor microenvironment, this work aims to generate more-durable and more-effective adoptive cellular immunotherapies for brain cancer in both adults and children.
Co-opting lactate signaling to enhance T cell therapy
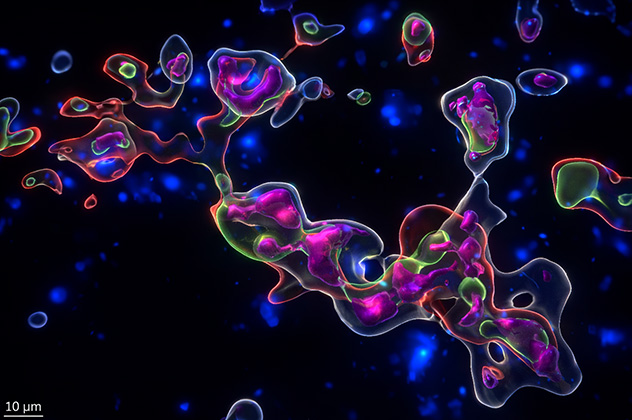 Immunometabolism at the tumor-immune interface
Immunometabolism at the tumor-immune interface
This image shows 3D confocal visualization of the glioblastoma microenvironment depicting direct metabolic exchange between tumor and immune cells. These spatially restricted interactions highlight why effective therapeutic strategies, including immunotherapies, must consider and leverage tumor and immune metabolic properties to achieve durable antitumor responses.
Glioblastoma is characterized by profound metabolic reprogramming consistent with the Warburg effect, resulting in high rates of aerobic glycolysis and excessive lactate production. Lactate accumulation within the tumor microenvironment contributes to immune suppression and impaired T-cell function, representing a major barrier to successful immunotherapy.
Building on this metabolic landscape, our laboratory is exploring innovative strategies to convert lactate from a tumor-derived liability into a therapeutic advantage.
This research project focuses on engineering immune cells with modified lactate-sensing receptors capable of translating lactate signals into pro-activation or costimulatory cues. By rewiring how immune cells interpret tumor-derived metabolic signals, we aim to overcome metabolic immune suppression and enhance the efficacy of adoptive immune cell therapies in glioblastoma.
Cross-cutting features of our research programs
Across all our research projects, our laboratory emphasizes:
- Use of patient-derived tumor models.
- Integration of single-cell and spatial profiling technologies.
- Mechanistic dissection of immunometabolic pathways.
- Strong alignment with translational and preclinical therapeutic development.